주메뉴
- About IBS 연구원소개
-
Research Centers
연구단소개
- Research Outcomes
- Mathematics
- Physics
- Center for Underground Physics
- Center for Theoretical Physics of the Universe (Particle Theory and Cosmology Group)
- Center for Theoretical Physics of the Universe (Cosmology, Gravity and Astroparticle Physics Group)
- Dark Matter Axion Group
- Center for Artificial Low Dimensional Electronic Systems
- Center for Quantum Nanoscience
- Center for Exotic Nuclear Studies
- Center for Van der Waals Quantum Solids
- Center for Relativistic Laser Science
- Center for Trapped Ion Quantum Science
- Chemistry
- Life Sciences
- Center for Memory and Glioscience (Cognitive Glioscience Group)
- Center for Memory and Glioscience (Learning and Memory Group)
- Center for Synaptic Brain Dysfunctions
- Center for RNA Research
- Center for Genomic Integrity
- Center for Vascular Research
- Center for Genome Engineering
- Center for Microbiome–Body–Brain Physiology
- Earth Science
- Interdisciplinary
- Center for Neuroscience Imaging Research (Neuro Technology Group)
- Center for Neuroscience Imaging Research (Cognitive and Computational Neuroscience Group)
- Center for Algorithmic and Robotized Synthesis
- Center for Nanomedicine
- Center for Biomolecular and Cellular Structure
- Center for 2D Quantum Heterostructures
- Institutes
- Korea Virus Research Institute
- News Center 뉴스 센터
- Career 인재초빙
- Living in Korea IBS School-UST
- IBS School 윤리경영


주메뉴
- About IBS
-
Research Centers
- Research Outcomes
- Mathematics
- Physics
- Center for Underground Physics
- Center for Theoretical Physics of the Universe (Particle Theory and Cosmology Group)
- Center for Theoretical Physics of the Universe (Cosmology, Gravity and Astroparticle Physics Group)
- Dark Matter Axion Group
- Center for Artificial Low Dimensional Electronic Systems
- Center for Quantum Nanoscience
- Center for Exotic Nuclear Studies
- Center for Van der Waals Quantum Solids
- Center for Relativistic Laser Science
- Center for Trapped Ion Quantum Science
- Chemistry
- Life Sciences
- Center for Memory and Glioscience (Cognitive Glioscience Group)
- Center for Memory and Glioscience (Learning and Memory Group)
- Center for Synaptic Brain Dysfunctions
- Center for RNA Research
- Center for Genomic Integrity
- Center for Vascular Research
- Center for Genome Engineering
- Center for Microbiome–Body–Brain Physiology
- Earth Science
- Interdisciplinary
- Center for Neuroscience Imaging Research (Neuro Technology Group)
- Center for Neuroscience Imaging Research (Cognitive and Computational Neuroscience Group)
- Center for Algorithmic and Robotized Synthesis
- Center for Nanomedicine
- Center for Biomolecular and Cellular Structure
- Center for 2D Quantum Heterostructures
- Institutes
- Korea Virus Research Institute
- News Center
- Career
- Living in Korea
- IBS School
News Center
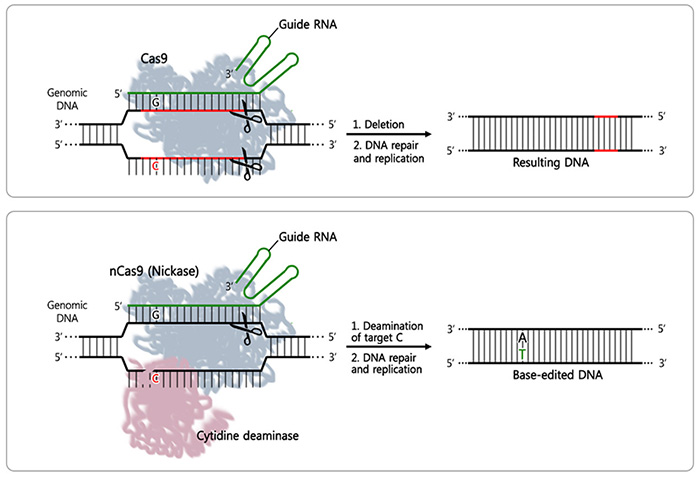
DNA Misspelling Correction Method is Very Accurate- IBS scientists prove that a gene editing technique used for substituting a single nucleotide in the genome is highly accurate - Researchers at the Center for Genomic Engineering, within the Institute for Basic Science (IBS) proved the accuracy of a recently developed gene editing method. This works as DNA scissors designed to identify and substitute just one nucleotide among the 3 billion nucleotides of our genome. "It is the first time that the accuracy of this base editor has been verified at the whole genome level," explains KIM Jin-Soo, leading author of this study. Published in Nature Biotechnology, this validation will help to expand the use of this method in the sectors of agriculture, livestock, and medicine, e.g. for gene therapy. Rapid progress in gene editing tools has caused a frenzy excitement in the biology community. The main protagonist of the current third-generation DNA scissors is CRISPR - a tool that is quicker and cheaper than its predecessors. By cutting out a small DNA sequence, CRISPR-Cas9 and CRISPR-Cpf1 are used to silence or reduce the expression of faulty genes. However, last year, a new base editor method that does not cause random DNA deletions and insertions, but instead replaces only one DNA base, attracted the biologists' attention. These types of gene corrections are critical as several diseases are caused by the misspelling of one of the four basic components of DNA; adenine (A), cytosine (C), guanine (G), and thymine (T). Single-nucleotide errors in DNA are referred to as point mutations. Examples of diseases caused by point mutations include: cystic fibrosis, sickle cell anemia, and color blindness. Unlike the existing third-generation DNA scissors, the base editor method consists of a variation of CRISPR-Cas9 (nCas9, nickase) fused with another enzyme called cytosine deaminase, which replaces the DNA component C with T. The scissors are directed to the correct position on the DNA by a guide RNA. However, up to now, it was not known whether the base editor was working only in the area of the faulty gene or if it was unnecessarily substituting Cs in other areas (off-target). Just one month after reporting the first successful base editing in animals in Nature Biotechnology to modify a single nucleotide in dystrophin and tyrosinase genes, the same team demonstrated the accuracy of this method at the genome scale.
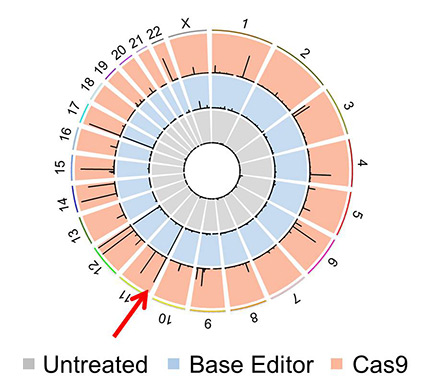
In order to identify the correctness of the gene editing for the entire genome, IBS researchers modified the error-checking technique, known as Digenome-seq, in order to adapt it to the base editor method. Digenome-seq was used and validated last year, when the team analyzed the accuracy of CRISPR-Cpf1 and Cas9. IBS researchers also improved the computer program (Digenome 2.0) to identify off-targets more comprehensively and compared different guide RNAs, to find the one that reduces malfunctions and increases specificity. Using this technique, the team demonstrated correctness of the base editor technique and they found it to be even more accurate than the current third-generation CRISPR-Cas9. The base editing technique induced C-to-T conversions in 1-67 sites in the human genome, while CRISPR-Cas9 caused cleavages in 30-241 sites, meaning that the base editor is making less off-target changes. "Therefore, it is expected that these base editors will be used as widely as the popular CRISPR technology," enthuses KIM.
Letizia Diamante Notes for editors - References - Media Contact - About the Institute for Basic Science (IBS) |
|||
Center for Genome EngineeringPublication Repository |
|||
|
|
| Next | |
|---|---|
| before |
- Content Manager
- Public Relations Team : Suh, William Insang 042-878-8137
- Last Update 2023-11-28 14:20